High-performance Computing with Amazon's X1 Instance
Eduardo Ariño de la Rubia2016-10-05 | 10 min read

We’re excited to announce support for Amazon’s X1 instances. Now in Domino, you can do data science on machines with 128 cores and 2TB of RAM — with one click:
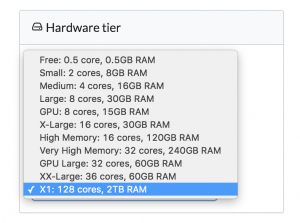
The X1 hardware tier is available in our cloud-hosted environment and can be made available to customers using Domino in their own VPCs.
Needless to say, with access to this unprecedented level of compute power, we had some fun. Read on for some of our reflections about doing data science with X1 instances.
Processing Power: Working with 128 Cores Under the Hood
Access to 128 cores on a single machine was nearly unheard of even just a few years ago, much less on a platform which could trivially be rented by the minute. Core counts at this scale were previously only the domain of distributed and HPC systems.
The ability to distribute a machine learning workload to 128 cores is a non-trivial problem, but two common techniques are (1) parallelizing the machine learning itself and (2) parallelizing fitting of the algorithm across multiple possible configurations (i.e., grid search).
Parallelizing grid search is fairly straightforward, and packages like scikit-learn and caret offer great solutions for this. Parallelization of a machine learning algorithm, however, is a challenging problem. There are a number of natural limitations to this approach, not least of which are the large-scale matrix operations at the core of many machine learning algorithms. These have natural bounds on the amount of parallelism that can be beneficial.
To explore these limits, I undertook a short and incomplete analysis of two modern machine learning toolkits, H2O and XGBoost for the task of fitting a GBM with 1,000 trees on the canonical airline dataset. I don’t undertake the task of validating the goodness of fit of the models generated. In this case, I’m simply interested in seeing how much parallelism these two packages are able to leverage when given a large number of cores.
Using H2O's R package version 3.10.0.6 and training on 100k rows of the airline dataset, the system was able to train a single model with 1,000 trees in 813 seconds. Full theoretical processor utilization would be 12,800%, that is, 100% utilization for each core. During training, processor utilization peaked at roughly 5,600%, implying 56 cores were in use.
Given the nature of the GBM algorithm, this limitation is understandable. There is an explicit limit on the amount of parallelism possible for training as determined by the shape of the input to the algorithm. It is also interesting to note that while peak memory usage of 46GB is high for GBM, it was still a very small percentage of total available RAM on the X1. Although H2O’s GBM algorithm provides excellent performance, it was not able to harness most of the processing power and memory available with an X1 instance.
When fitting multiple models and attempting to search a large hyperparameter space, the power of the X1 instance type and H2O’s Grid tools show value. Using a modified version of H2O’s Grid Search example H2O’s package was able to utilize almost all 128 cores!
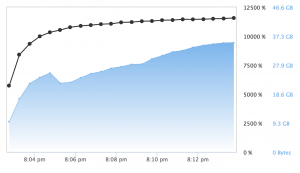
While generating any single individual model with H2O’s grid search was limited by the GBM algorithm, the ability to search a vast hyperparameter space and take advantage of the X1 to find the optimal hyperparameters for a GBM should not be undervalued as this is an often time-consuming and tedious task.
Results were better when training a similar single GBM model using the XGBoost package for R. XGBoost was able to leverage nearly 9300% of the system’s available processor power, or 93 cores. While this still leaves some number of processors under-utilized, XGBoost was able to train a GBM with 1000 trees in 117 seconds, under 1/7th of the time needed by H2O’s GBM.
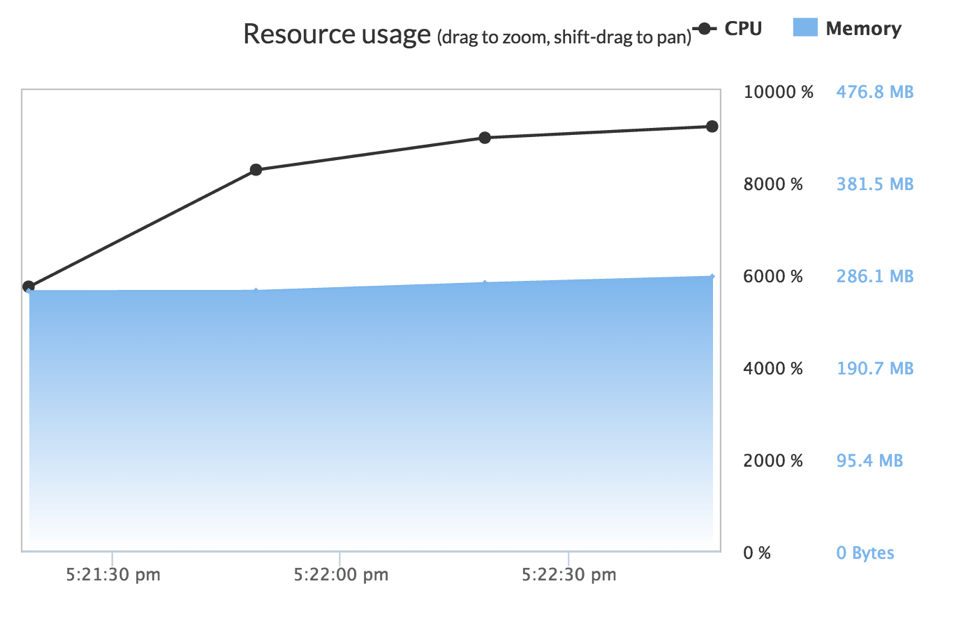
XGBoost’s author, Tianqi Chen has spoken extensively about the many tricks and approaches that he has leveraged in order to make it take advantage of massive parallelism. XGBoost’s singular focus on providing a high-performance GBM implementation shows: Its finely-tuned approach is able to take considerable advantage of the hardware of the X1, training models in less than ⅓ of the time taken when using the largest 36-core hardware option previously available on the Domino platform.
Also of note is that XGBoost was incredibly efficient with memory usage, using just 286 megabytes of memory training on the same dataset as H2O. This is roughly 0.5% of the memory allocated by the H2O implementation and its runtime. Though it was not an issue on the X1 instances, given their vast amount of addressable memory, it was an interesting finding.
Enough Processing Power to Encroach on Distributed Systems
Leveraging the AWS X1 through Domino provides data scientists with processing power far beyond what is commonly available from a single machine. While all of the processing power isn’t immediately and simply accessible and requires some intelligent application, being able to leverage 90+ cores when training a GBM inside of XGBoost, for example, pushes the boundary of what problems require a distributed system.
Problems that would previously have needed a distributed system such as Apache Spark, and required a re-implementation and re-training on potentially unfamiliar tools, can now be solved with relative simplicity. The ability to train GBM’s and do massive parallel grid search using tools that data scientists already know and love will force many distributed system adherents to question at what scale they really need to pay the penalty of a distributed system. When one can train 1,000 trees on 1 million rows of data in 3 minutes, the bar has been moved considerably. It should be noted that this is not a benchmark of model performance. If the reader is interested in a deeper analysis of the performance of models, this work by Szilard Pafka is highly regarded and compares the libraries used in this post and many more.
Memory: Working with 2TB of RAM Has its Challenges
2TB of RAM is a preposterous amount. A back of the envelope calculation shows that on the X1 instance it is possible to load 42 billion rows of data in RAM for real-time interactive exploration. Similarly, when thinking about this in terms of graph data, one could viably load a graph which is over 8,000 times the size of the LiveJournal graph dataset.
The challenge at this scale is rather surprising: how does one even load 2TB of data into RAM, and how long does that take? Leveraging data from a previous blog post, we discovered that the data.table fread is a very fast way of reading data from disk. Some quick benchmarking showed that reading at 70MB/s is a reasonable expectation for the single-threaded read_csv from the readr package. This would mean a total read time of close to 8 hours for 2TB of data! While there are a number of multi-core CSV parsers available, users would still likely need to wait up to an hour for the data load to complete.
This is a rather unexpected aspect of dealing with such large datasets.
Conclusion
Leveraging the X1 instance from Domino provides data scientists a true “supercomputer in a box” experience. Problems that previously required porting to a complex distributed computing environment such as Apache Spark and the use of unfamiliar languages such a Scala, can now be handled by the tools data scientists know and love: R and Python.
Tasks which are traditionally parallelizable, such as grid search or Monte Carlo simulations, can trivially take advantage of the significant number of available cores. While many modern machine learning libraries do take advantage of parallelism, they won’t immediately take advantage of all cores, and will require some work to fully harness the power of the X1.
Having 2TB of RAM available allows an entirely new class of “whole-dataset” analysis which previously would have required expensive trips to disk and constant reckoning with the logarithmic latency penalty of access to longer term storage. However, as always there is no free lunch: Rapidly reading data from disk into RAM still presents an interesting challenge.
We're excited by the opportunity to enable your analysis on this scale of hardware, and can’t wait to see what kinds of new analysis and new models this platform allows you to create. X1 instances are now available in Domino’s cloud production environment, and we will gladly work with any interested VPC customers in getting X1 instances provisioned into their environment.
Curious about what other algorithms can benefit from leveraging the powers of X1? I had some fun in Part II.
Banner image titled "Bookends" by Nic McPhee. Licensed under CC BY 2.0
Eduardo Ariño de la Rubia is a lifelong technologist with a passion for data science who thrives on effectively communicating data-driven insights throughout an organization. A student of negotiation, conflict resolution, and peace building, Ed is focused on building tools that help humans work with humans to create insights for humans.
Subscribe to the Domino Newsletter
Receive data science tips and tutorials from leading Data Science leaders, right to your inbox.
By submitting this form you agree to receive communications from Domino related to products and services in accordance with Domino's privacy policy and may opt-out at anytime.



